Part 1 of 6 in the SCARCE-CXR series
1.1 My First Failure
When I was just starting out with machine learning, I tried to build a chest X-ray classifier. I found a small binary dataset of around 600 labeled X-rays, cancer or not cancer, threw a RandomForest at it, got 55% accuracy, and convinced myself I was getting somewhere.
I wasn't.
The confusion came from not thinking carefully about what 55% means on a balanced dataset. Random chance is 50%, so 5% above coin flip is actually not significant at all. There was nothing separating that margin from a scanner brightness difference between batches (I'm pretty sure that's what it was).
The deeper problem was the model itself. RandomForest operates on flattened pixels. A downscaled 512×512 chest X-ray has 512×512=262,144 input dimensions with spacial pathological signal: the shape of a nodule, its texture relative to surrounding lung parenchyma, and whether margins are irregular or smooth. There was no way to learn that an irregular opacity in the upper lobe signals malignancy with spatial relationships flattened into a 1D vector.
It needed convolutional structure it didn't have, trained on volume it also didn't have.
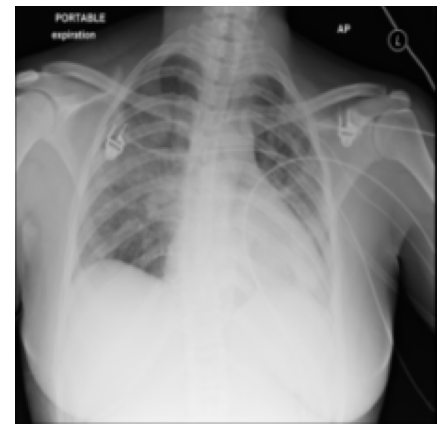
1.2 Throwback to 2020
I wasn't the only one making this mistake. Around the same time, the entire ML community tried to build COVID classifiers from scratch. It was a larger-scale version of the same failure.
The data situation was dire, but researcher Joseph Paul Cohen did something heroic and scraped chest X-rays from published PDF medical journals to build a public repo. Soon there were around 100-200 positive COVID X-ray images in existence. To train a binary classifier you need negatives, so researchers grabbed the largest available chest X-ray dataset: the Kermany dataset on Kaggle. Fast, free, accessible. The fatal flaw nobody caught until later: the Kermany dataset was collected from the Guangzhou Women and Children's Medical Center. It's almost entirely pediatric patients, ages 1-5. The COVID images were adults.
Thousands of papers were published claiming 95–98% accuracy. DeGrave et al. audited the top-performing ones with saliency maps. The models were not looking at the lungs. They were detecting hospital-specific "L/R" text marker fonts, JPEG compression artifacts from the PDF scraping, and patient positioning: sick COVID patients were supine in ICU beds, healthy controls were standing. Roberts et al. reviewed 415 published COVID detection models and found zero fit for clinical use. Maguolo and Nanni blacked out the lungs entirely and still got near-perfect "COVID detection" from border artifacts and padding.
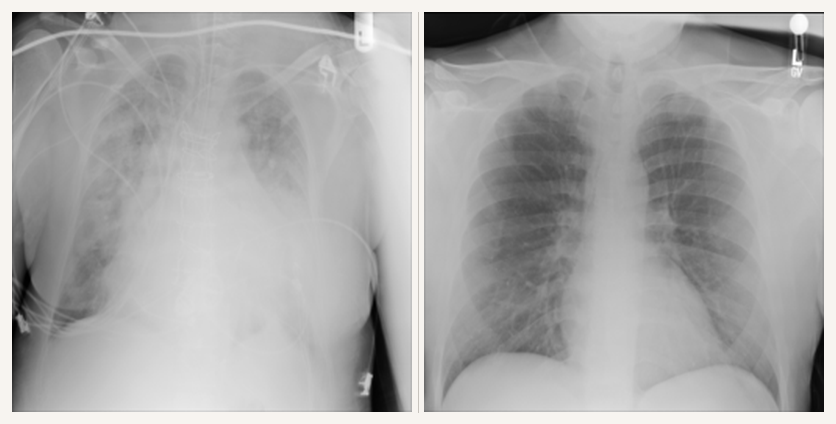
This is what supervised learning does on small datasets: it overfits and finds shortcuts. Learning the texture of ground-glass opacities is computationally hard. Learning that COVID images have fuzzier JPEG compression or a specific text font is computationally easy. The optimizer takes the easy path every time, just as it was designed to. Imaging was never going to replace swabbing for diagnosis, but it could have helped in flagging severity, prioritizing ICU beds, catching deterioration early. That window closed without a working model.
You cannot learn the complex anatomical structure of a human chest and a novel disease at the exact same time. The problem wasn't the classifiers. It was the foundation.
1.3 The Real Problem
A couple of years later I came back to medical imaging with a clearer understanding of what the actual problem is.
It's not "can we train a chest X-ray classifier?" That problem is mostly solved. NIH has 112,000 labeled chest X-rays. Stanford's CheXpert has 224,000. For common diseases (pneumonia, pleural effusion, cardiomegaly), you can train a ResNet50 supervised from scratch and do reasonably well. There's enough labeled data.
The problem is this: what do you do when you only have 23 labeled examples?
Think about what that scenario looks like in practice. A small regional hospital has been flagging an unusual pattern of interstitial lung disease in a handful of patients. They have 23 annotated X-rays. There is no large public dataset for this finding. As the COVID models proved, you cannot train a convolutional network from scratch on 23 images. The gradients are noisy, the model overfits immediately, and what you end up with is worse than a simple threshold on pixel statistics. Direct supervised training is not the answer.
But there is an alternative: Self-Supervised Learning (SSL).
Instead of treating the 112,000 images in the NIH dataset as a supervised classification task, we can disregard the labels completely and use them to teach a backbone what chest X-ray structures look like. It learns what a ribcage looks like, where the lung is positioned, what normal scanner variation looks like. This directly addresses both failure modes:
- Overfitting is neutralized: the backbone trains on 112,000 images with no labels to memorize. By the time it sees your 23 labeled examples, the heavy lifting of representation learning is already done.
- Shortcut learning is blocked: models cheat because they are searching for the easiest shortcuts (an "L" marker) that correlates with a disease label. By pretraining without labels, you remove the answer key. The model is forced to learn the holistic structure of the chest itself.
Then, when you hand the pretrained backbone 23 labeled examples of something rare, the only work left is identifying the new pathological pattern on top of a foundation that already understands chests.
That is the bet. Part 2 explains how it's built and where it breaks.
1.4 Sourcing Data
I pretrained on NIH ChestX-ray14: 112,000 frontal chest X-rays from the NIH Clinical Center, free on Kaggle. I threw away all the disease labels and used it purely as unlabeled image data. Training uses PadChest: a Spanish hospital dataset with 160,000 X-rays and a long tail of rare findings with almost no labeled examples elsewhere.
Using two datasets from different countries was deliberate. The 2020 COVID models failed precisely because they trained and tested within the same distribution. If my backbone is learning NIH scanner quirks, those shortcuts won't transfer to a Spanish hospital machine, and the AUC will drop. Cross-hospital performance is the test. If it holds, the model learned anatomy, not acquisition artifacts.
Everything here is free to reproduce. NIH is on Kaggle. PadChest needs an email approval. All SSL methods ran on a single L4 GPU using Google Cloud free trial credits.
Here's how I built it.